My dissertation, “Interpretable machine learning in plant genomes” (available here, focused on developing and interpreting machine learning models to study complex biological systems in plants. This work focused on three areas:
GENOMIC PREDICTION GENE REGULATION OPINIONS ON BIOTECHNOLOGY
GENOMIC SELECTION
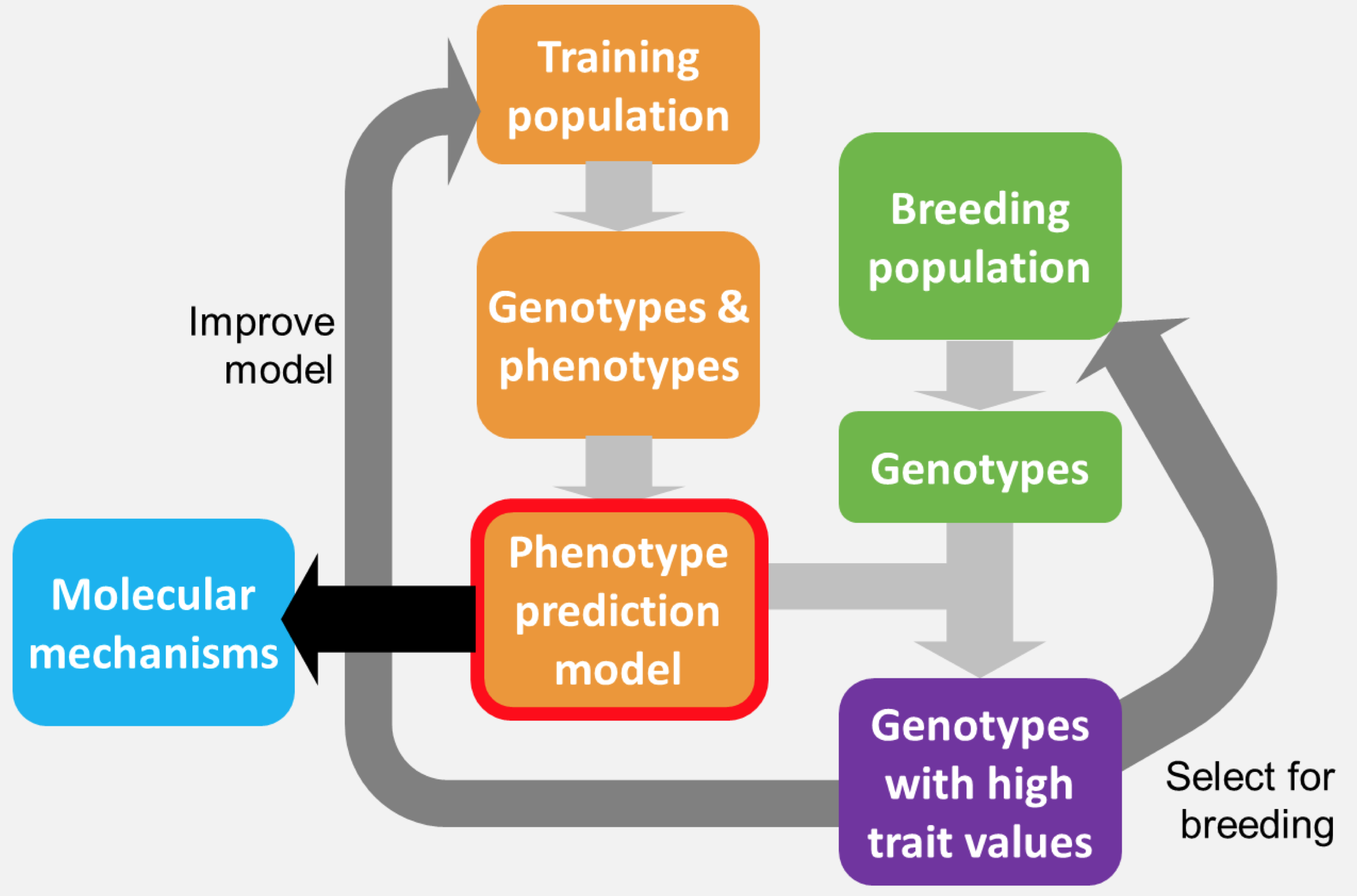
The ability to predict complex traits from genotypes is a grand challenge in biology and is changing crop and livestock breeding. Genomic Prediction is the prediction of traits from genetic sequence. It relies of identifying of tens, to hundreds, to thousands of genetic variants that are predictive of complex traits, such as crop yield or disease seceptability. whole-genome DNA sequence was originally proposed by Meuwissen et al. in 2001 as a solution to the limitations of Marker-Assisted Selection (MAS). GP is particularly well-suited for the prediction of quantitative traits, such as yield or drought resistance, that are controlled by many small-effect alleles. Since 2001, there has been an explosion in GP algorithms available, that differ in how they account for non-linear interactions, population structure, differences in the effect size at each marker, and epistatic interactions between markers. I
Most recently, deep learning approaches have been used to predict traits from genomic data. Briefly, deep learning refers to machine learning approaches that perform layers of transformations on features to create abstraction features, known as hidden layers, which are used for the ultimate predictions. An ongoing effort is to benchmark such deep learning models against more tradiational GP algorithms across a broad range of species and traits.
GENE REGULATION
I am fascinated by the complexity of how crop plants respond to environmental stresses, particularly stresses disrupting agricultural systems due to climate change. Plants are diverse in their response to stress, yet across species these pathways are tightly regulated and largely interwoven. By interwoven, I mean that many of the components that regulate response function in response to more than one kind of stress. This “crosstalk” between pathways makes it difficult for us to predict how plants will respond to multiple stresses based on the kind of single stress studies commonly done in labs.
In nature, plants are rarely exposed to just one stress at a time, so understanding how these response pathways talk to each other is critical to understand how plants will respond to climate change. The first project I’m tackling during my Ph.D. is to uncover the cis-regulatory code of plant response to combined stress using multi-dimensional data integration and machine learning.
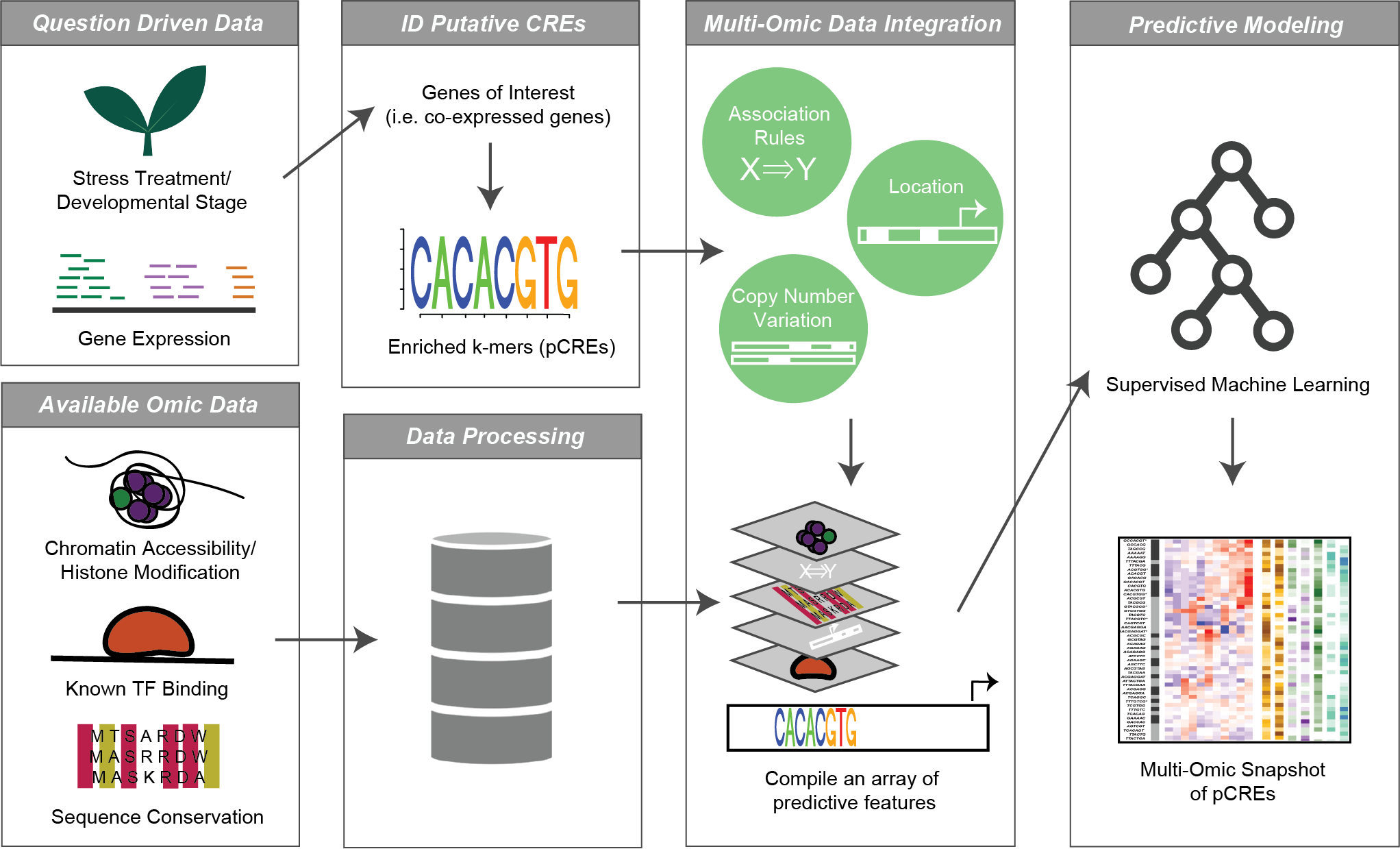
In a nutshell, I identify short sequences in gene promoter regions that are common between genes that are co-expressed under single and combined stress scenarios. Then I use information about those sequences, like their overlap with known TF binding, their chromatin accessibility, proximity to histone markers, sequence conservation, and more to assess how likely they are to be true regulatory elements. Finally, I build predictive models using machine learning to determine how well these putative regulatory elements can be used to predict a gene’s response to the stress conditions.
PUBLIC OPINION OF BIOTECHNOLOGY

Research on public views of biotechnology has centered on genetically modified (GM) foods. However, as the breadth of biotechnology applications grows, a better understanding of public concerns about non-agricultural biotechnology products is needed in order to develop proactive strategies to address these concerns. As an Environmental Science Policy Program (ESPP) fellow at Michigan State University I identified this a major problem facing those working to develop biotechnology policy. Therefore, as my capstone project I conduced a survey to explore the perceived benefits and risks associated with five biotechnology products and how those perceptions translate into public opinion about the use and regulation of biotechnology. READ THE PUBLICATION