I am fascinated by the complexity of how crop plants respond to environmental stresses, particularly stresses disrupting agricultural systems due to climate change. Plants are diverse in their response to stress, yet across species these pathways are tightly regulated and largely interwoven. By interwoven, I mean that many of the components that regulate response function in response to more than one kind of stress. This “crosstalk” between pathways makes it difficult for us to predict how plants will respond to multiple stresses based on the kind of single stress studies commonly done in labs.
In nature, plants are rarely exposed to just one stress at a time, so understanding how these response pathways talk to each other is critical to understand how plants will respond to climate change. The first project I’m tackling during my Ph.D. is to uncover the cis-regulatory code of plant response to combined stress using multi-dimensional data integration and machine learning.
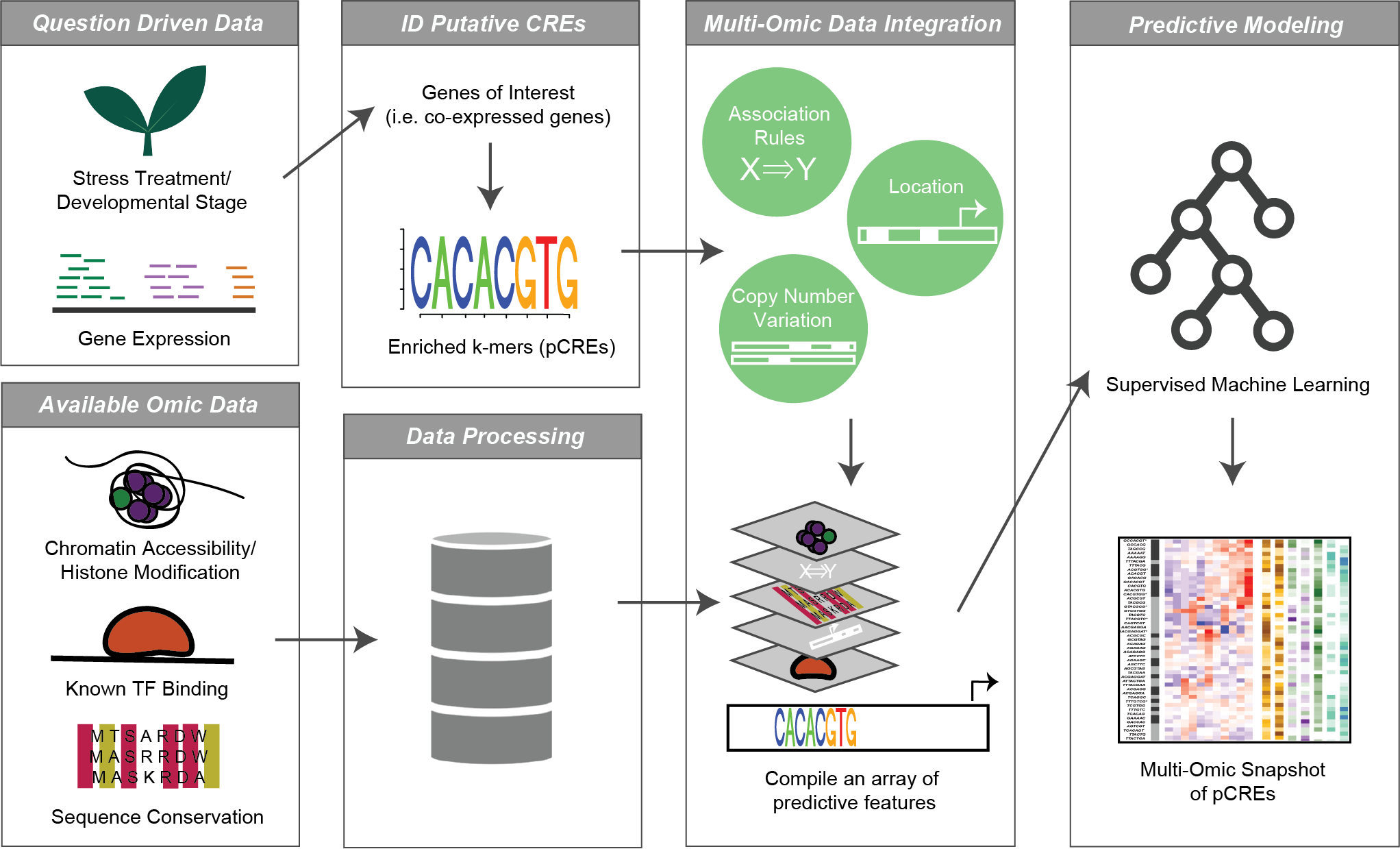
In a nutshell, I identify short sequences in gene promoter regions that are common between genes that are co-expressed under single and combined stress scenarios. Then I use information about those sequences, like their overlap with known TF binding, their chromatin accessibility, proximity to histone markers, sequence conservation, and more to assess how likely they are to be true regulatory elements. Finally, I build predictive models using machine learning to determine how well these putative regulatory elements can be used to predict a gene’s response to the stress conditions.