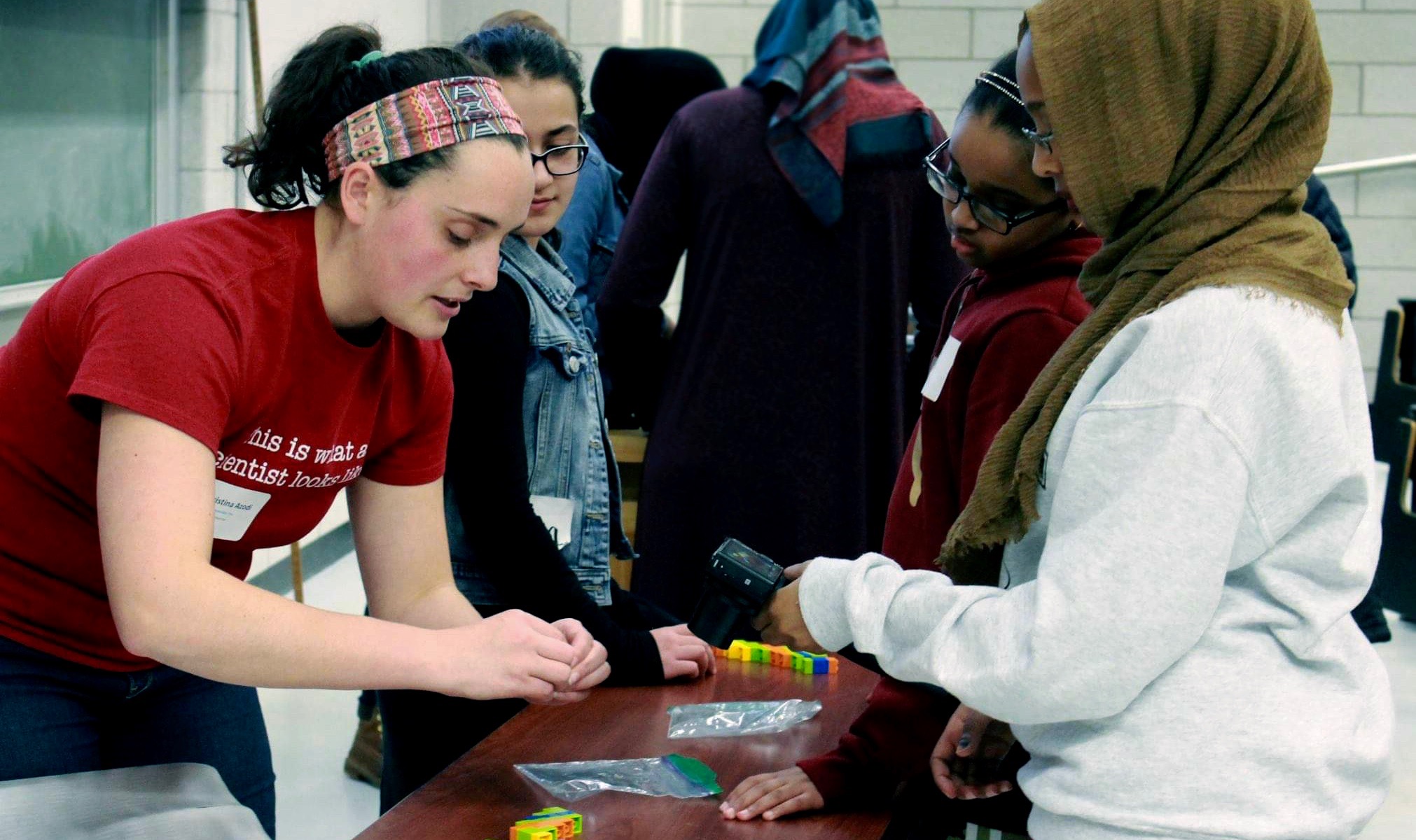

Engaging young people in computational research can be challenging. To help get kids excited about programming and biology, I created two hands-on and computer-free activities for teaching skills in computer science and bioinformatics. The activities can be adjusted to suite a range of age groups. I would love to hear from your if you try any of these out!
Code Like a Girl
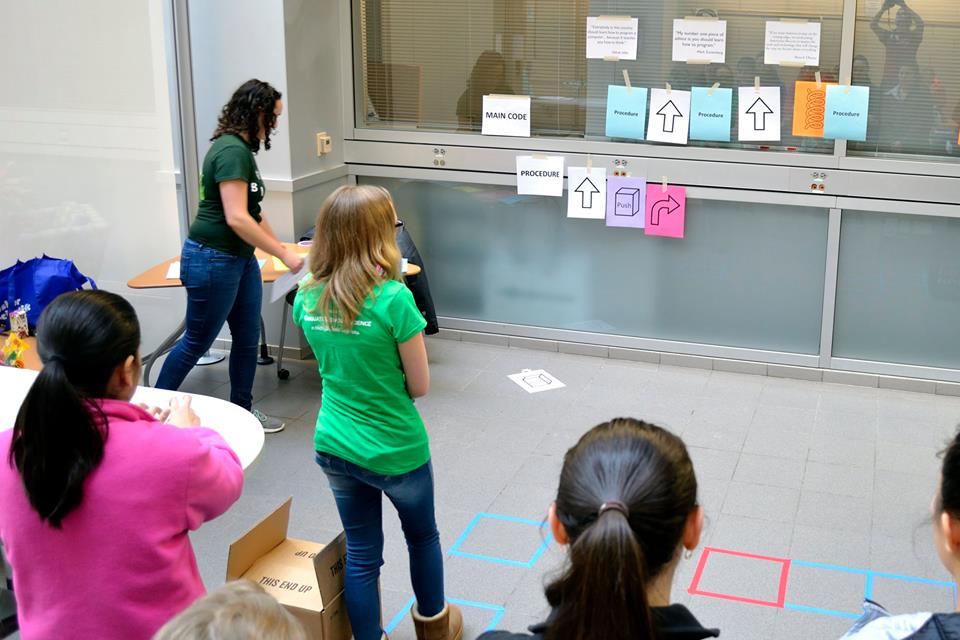

Learn to think like a computer scientists by writing a program that will get you and your friends around obstacles to make it through a treacherous maze!
While this activity was originally developed for middle school age girls for “Girl’s Math and Science Day” @ Michigan State University, hosted by the MSU Graduate Women in Science, it is suitable for all elementary school age students. For students much younger than middle school, simpler mazes may need to be designed.
The activity is based loosely off LightBot, where you have a “robot” (i.e. one of the students) who you are helping through a maze by designing a “program” of step-by-step instructions to make it through the maze. Things get tricky when you introduce functions and limit the number of moves.
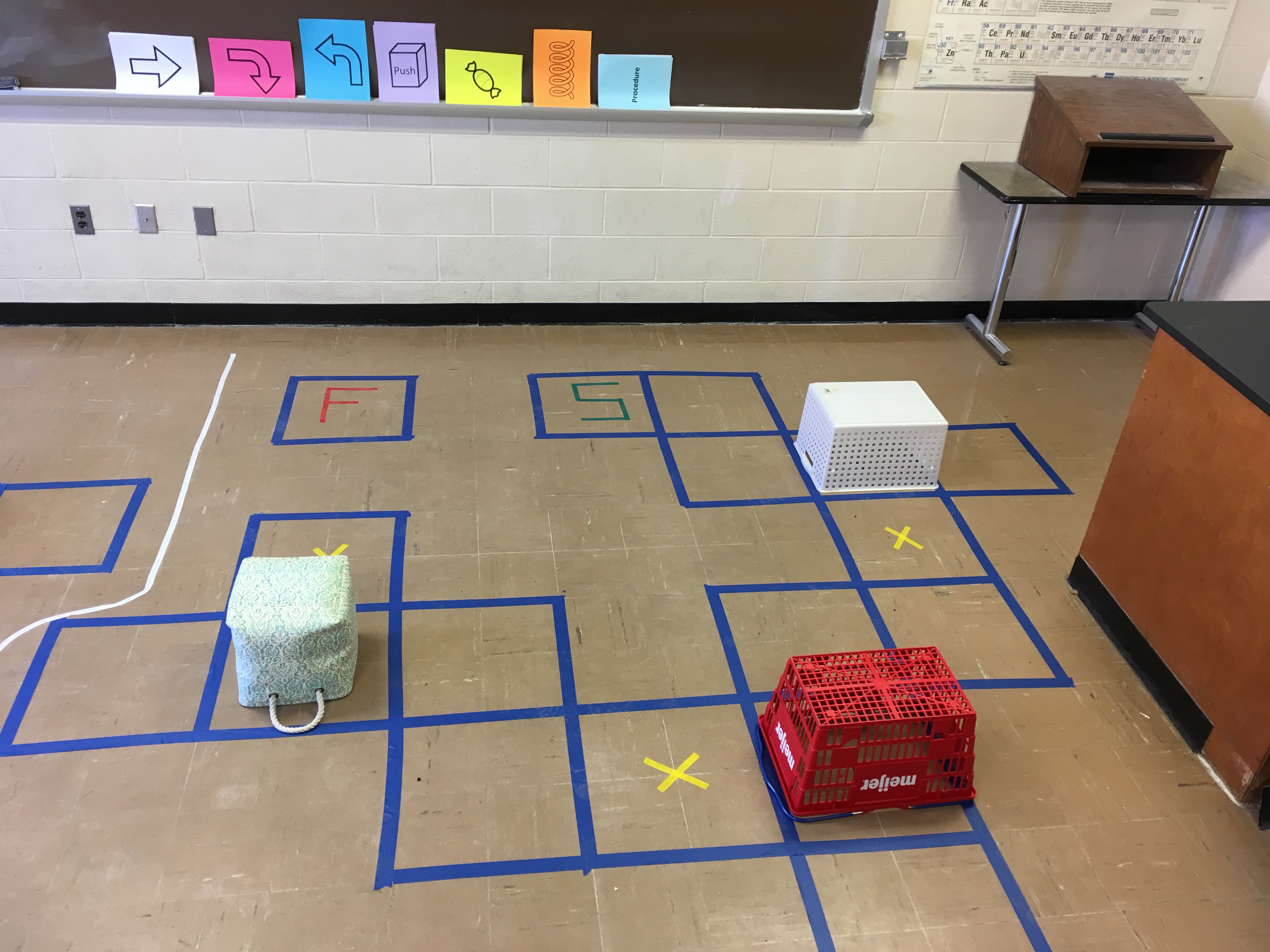

Find all of the puzzles and the code cards here!
Step 1. Explain the game with puzzle 1
Explain what the code cards do (i.e. move forward, turn left, turn right, jump, push the box, pick up the candy, etc). Then have the girls write a code to solve puzzle one using as many steps as they want. I like having a strip of tape on the cards and taping them to a black/white board (which is nice cause you can highlight the repeating patterns later).
Step 2. Writing functions
They probably figure out the game pretty quickly in step one, and if they had unlimited code cards it would be easy for them to solve any puzzle. But here is where you introduce functions. Discuss how computer programs are good for automating repetitive tasks. Then ask them to look for “repetitive” patterns in their code, and move the repetitive part to a function. Then they can run the code by calling their function when that pattern is needed, drastically reducing the number of code cards needed to guide their friend through the maize.
Step 3. Identifying patterns
Move on to puzzle 2. The goal of puzzle two is to make it from start to finish and pick up 3 candies on the way. There are 3 candies physically close to them and a few more that seem farther away. The trick here is that they don’t have enough code cards to get all of the close by candies, but instead have to write a function that allows them to get to the farther away candies.
Step 4. Challenge mode
Puzzle 3 is a lot of fun because there are a few solutions, but they all involve writing functions. The patterns in this puzzle are also less easy to spot, so it’s best to do this one after they’ve completed puzzle 2. If you’re working in a 30 minute time frame, I would suggest going through steps 1-2 together, then splitting into two groups, group one does puzzle 2 and group two does puzzle 3. Then at the end the groups can do a show-and-tell explaining what they had to do to the other group, talking about why it was difficult, and how they solved it.
Again, all of the puzzles and code cards are available here. If you have questions about how to implement this at your own science outreach event please find me on twitter @cbazodi. I would also be happy to add new puzzles to the document so if you design something new let me know!
Assemble This!
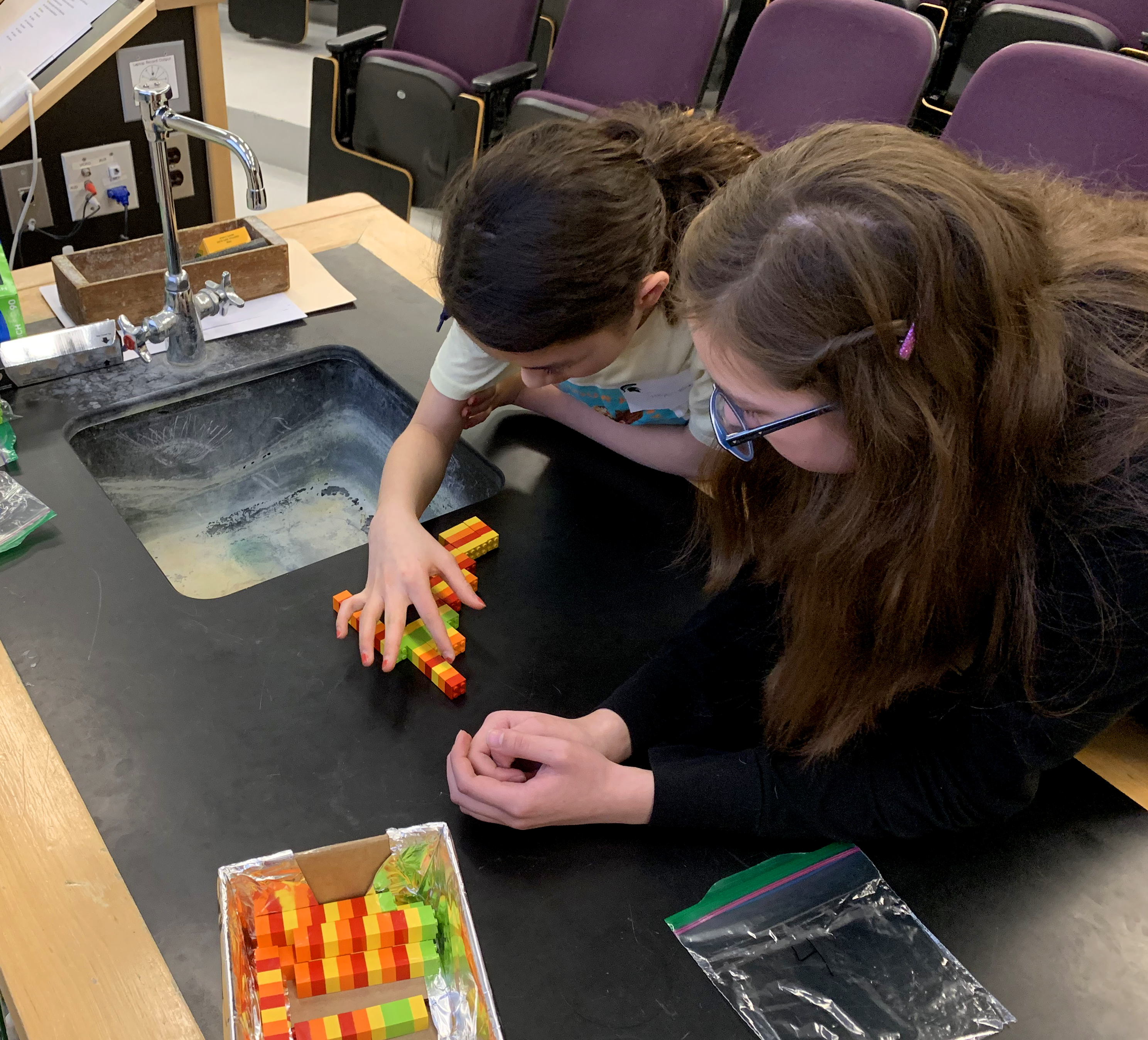

Using Legos as your DNA, test your skills at assembling genes while learning about human and plant genomes and about how scientists use computers to assemble whole genomes!
This activity was developed for middle school age girls for “Girl’s Math and Science Day” @ Michigan State University, hosted by the MSU Graduate Women in Science, it is suitable for all elementary school age students and can be adapted for younger or older students by developing easier or more challenging puzzles.
Before jumping into this hands on activity, we talked about what the girls knew about DNA and then gave a short presentation on genomes, whole genome sequencing, and why genome assembly is so challenging.
Finally, we introduced the rules of the activity:
- No altering the k-mers (no sticking them together and no taking them apart)
- Each part of the assembly will be supported by at k-mers with a 2 Lego overlap.
- All k-mers will face the same direction (i.e. the Lego dots point the same way).
Vocabulary: a k-mer is a short Lego fragment (4 Legos long) used to generate the assembly (all k-mers lined up by overlaps that reveals the final sequence).
The activity consisted of four, progressively more difficult puzzles. Each one took a pair of middle school age students between 5-10 minutes to solve. The first puzzle has some large runs of the same color, making it easier to assemble than the second puzzle (see pictures below). The biggest challenging was developing good puzzle solving skills, like assembling the pieces with the most overlap (i.e. greatest support) first and learning that because two k-mers overlapped didn’t necessarily mean they aligned with each other in the final puzzle.
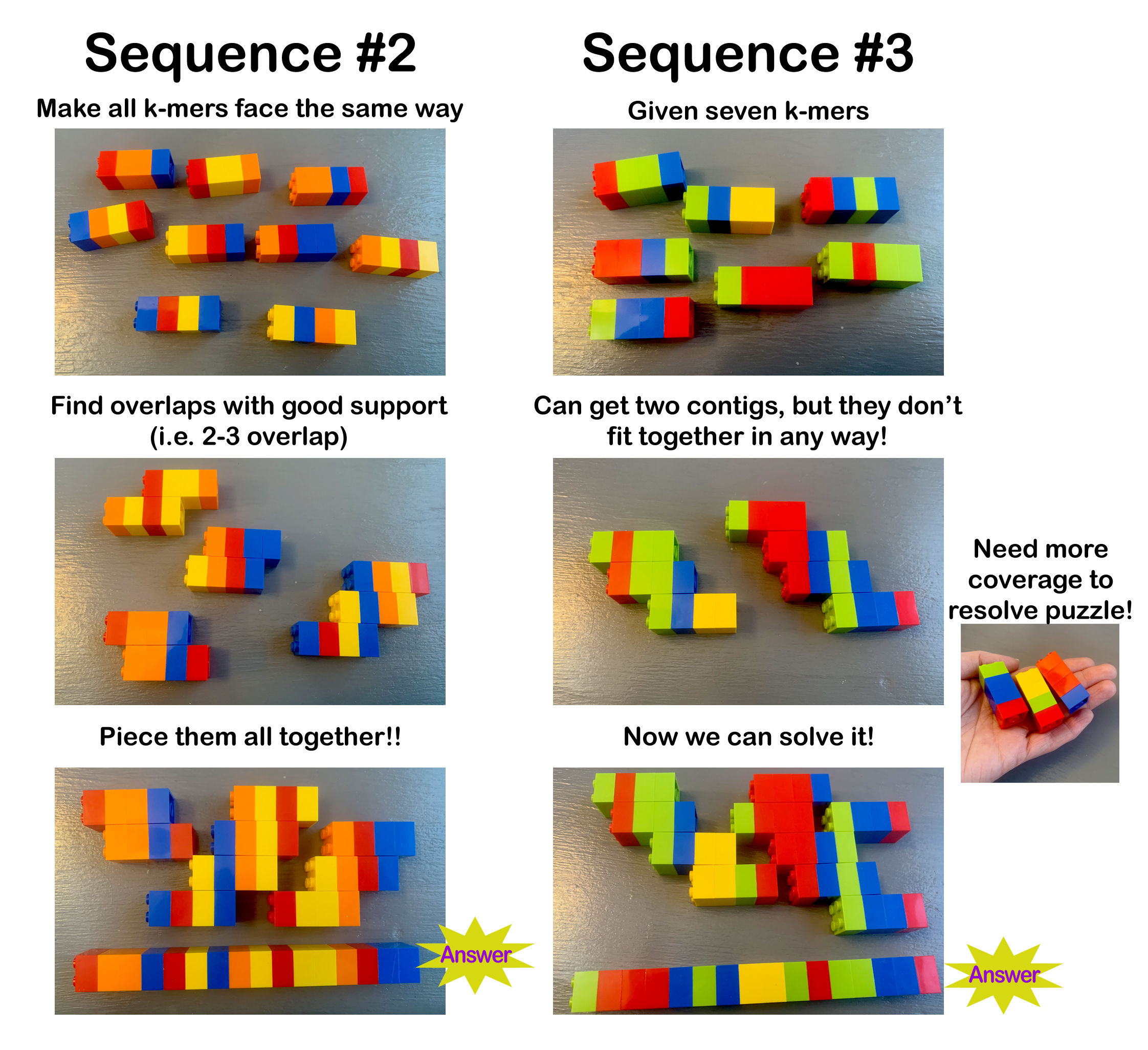
The next two puzzles were more challenging and were designed to teach the concepts of gap filling and repetitive sequence assembly. For puzzle 3, the students were given enough k-mers to resolve two separate parts of the sequence (i.e. contigs) but those parts could not fit together, leaving a gap. After letting them struggle with this problem for a few minutes, we stopped and talked about how gaps exist in the human genome and that one way to reduce the number of gaps is to get more sequencing done in order to have enough sequence support to fill a gap. To drive this lesson home, we provided each group with three additional k-mers that allowed them to fill the gap and connect the two short sequences. For puzzle 4, the students were given highly repetitive k-mers and were not told how long the sequence was supposed to be, after all, this is typically not know ahead of time. The students then quickly assembled short sequences with very uneven k-mer coverage. We then revealed that the answer was 17 Legos long! A few groups were able to spread out the k-mers to make their solution the correct length, but most struggled. At this point, we stopped and talked about how new sequencing technologies (e.g. long read sequencing) were allowing scientists to better resolve these repetitive regions. To drive this lesson home, we then provided each group with two long read k-mers that allowed them to resolve the 17-Lego long sequence.
Find the other puzzles and a break down of how many Legos you will need here!